Lecture 1: Basics
Course objectives, schedule, assignments · Introduction to R · Live coding

Helmholtzstraße 22, Room 205
andre.guettler@uni-ulm.de
+49 731 50 31 030

Helmholtzstraße 22, Room 203
oliver.padmaperuma@uni-ulm.de
+49 731 50 31 036
1.1 Course objectives
- 1.1 Course objectives
- 1.2 Introduction to R
- 1.3 Live Coding Session 1
- 1.4 Conclusion of Lecture 1
- Welcome to
- Course Objective
- Course at a glance
- Assignments / Exams
Welcome to Research in Finance
- Register for “exam” 13337 in campusonline by 30 November 2025. The registration is what binds you to the course requirements; without it you cannot submit. If you are registered but don’t submit, you receive a fail grade (5.0).
- Ask questions during or right after each session — that is the preferred channel.
- Admin / studies / exam-eligibility questions go to the registrar’s office (Studiensekretariat) at studiensekretariat@uni-ulm.de.
- Course-content questions outside class: email oliver.padmaperuma@uni-ulm.de, CC andre.guettler@uni-ulm.de.
- We also recommend the student advisory service.
Course Objective
Scope
We will:
- Prepare Master students for their empirical thesis
- Hands-on R intro for data management, visualization, cleaning, basic modelling
- Writing tips for theses, including LaTeX & Overleaf
- Referee reviews on research presentations for empirical critique skills
We will NOT:
- Deep dive into advanced stats or ML methods
- Specific finance topics (asset pricing, etc.)
- Full thesis writing / research design training
Approach
Part I — Learn the Basics
- Hands-on R intro: a widely used language for statistical computing
- Manage, visualize and clean data; run and interpret statistical models
- Solve a real empirical problem set in R, in groups
Part II — Apply your learnings
- Mandatory participation in the institute’s Brown Bag Seminar
- Two assignments (group work and individual referee report) — see Assignments / Exams
Course at a glance
Basics
Week 1
29.10.2025
Course objectives, schedule, assignments · Introduction to R · Live coding
- Course objectives, schedule and assignments
- Introduction to R and RStudio
- Live coding: variables, vectors, matrices, data frames, lists, functions, loops
- Data import and export
Data Handling & Visualization
Week 2
05.11.2025
API access, merging, cleansing, transforming and visualising financial data in R · Introduction to Overleaf
- API access (Nasdaq Data Link / Quandl, FRED, Yahoo, Coingecko, Polygon)
- Import and cleanse: read_csv, mutate, types
- Merge and append data (merge, bind_rows)
- Filter and mutate (dplyr): subset rows, derive variables
- Group by and summarise
- Pivot wide / long
- Data visualization with ggplot2 (six-step pipeline)
- Introduction to LaTeX and Overleaf
Statistical Analysis
Week 3
12.11.2025
Descriptive · inferential · modelling — applied in R
- Descriptive statistics in R
- Correlation matrix and Pearson correlation test
- t-Test and Wilcoxon test
- Shapiro-Wilk and Kolmogorov-Smirnov tests
- Linear regression with fixed effects
- Clustered standard errors
- Exporting regression tables with stargazer
- Discussion of Assignment I (Problem Set)
Academic Publishing & Refereeing
Week 4
19.11.2025
What makes a great empirical paper · publication process · how to write a referee report
- What makes a good empirical paper (contribution, identification, write-up)
- The publication process step by step
- Top finance and economics journals
- Bad outcome vs revise & resubmit
- Referee Reports — summary, major issues, minor issues
- Referee checklist (question, identification, data, econometrics, results)
- Discussion of Assignment II (Referee Report)
Brown Bag Seminar
Week 13
20.01.2026
Engage with doctoral research and prepare your referee report
- Doctoral research presentations
- Apply empirical / writing tips for the referee report
- Group discussion and Q&A
Assignments / Exams
Assignment I — Problem Set 50% of your grade
Documented .R script + PDF write-up (Overleaf)
Group of up to 5.
Submit by emailing oliver.padmaperuma@uni-ulm.de, CC andre.guettler@uni-ulm.de. Subject pattern: Research in Finance_assignment-1-problem-set_surname1_surname2_…
19 January 2026
Assignment II — Referee Report 50% of your grade
2.5–3 page referee report on a Brown-Bag presentation
Group of up to 5.
Submit by emailing oliver.padmaperuma@uni-ulm.de, CC andre.guettler@uni-ulm.de. Subject pattern: Research in Finance_assignment-2-referee-report_surname1_surname2_…
3 February 2026
1.2 Introduction to R
- 1.1 Course objectives
- 1.2 Introduction to R
- 1.3 Live Coding Session 1
- 1.4 Conclusion of Lecture 1
- Survey — let’s get to know you
- Why learn R?
- What are R and RStudio?
- Installing R and RStudio
- RStudio interface
- Writing code in RStudio
- Setting the working directory
- Getting started — running code
- What are packages?
- Installing and loading packages
Survey — let’s get to know you
Open the Mentimeter link shared in class.
Why learn R?
- It’s free and open-source.
- A powerful tool for data analysis and graphics.
- R code is great for reproducibility and shareability — submit your script and anyone can re-run your results.
- A large, active community continuously extends R via packages.
- Widely used in academia and industry — especially in finance, economics, and data science.

What are R and RStudio?
- R is a programming language for statistical computing, data analytics, and scientific research. It is one of the most widely used languages by statisticians and researchers to manage, manipulate, analyze, and visualize data.
- RStudio is an integrated development environment (IDE) for R that makes interaction with R easier — code completion, debugging, project management, plot viewing.
- In 2022 RStudio was rebranded Posit to signal its move toward language-agnostic tooling.
In order to use RStudio, you need to have R installed. RStudio is just the interface. R does the actual computing.
Installing R and RStudio
- Go to https://cran.r-project.org/ and click Download R for Windows / macOS.
- Click base subdirectory.
- Click Download R-x.x.x for Windows and run the
.exe. Accept defaults. - Go to https://posit.co/download/rstudio-desktop and click Download RStudio.
- Run the RStudio installer.
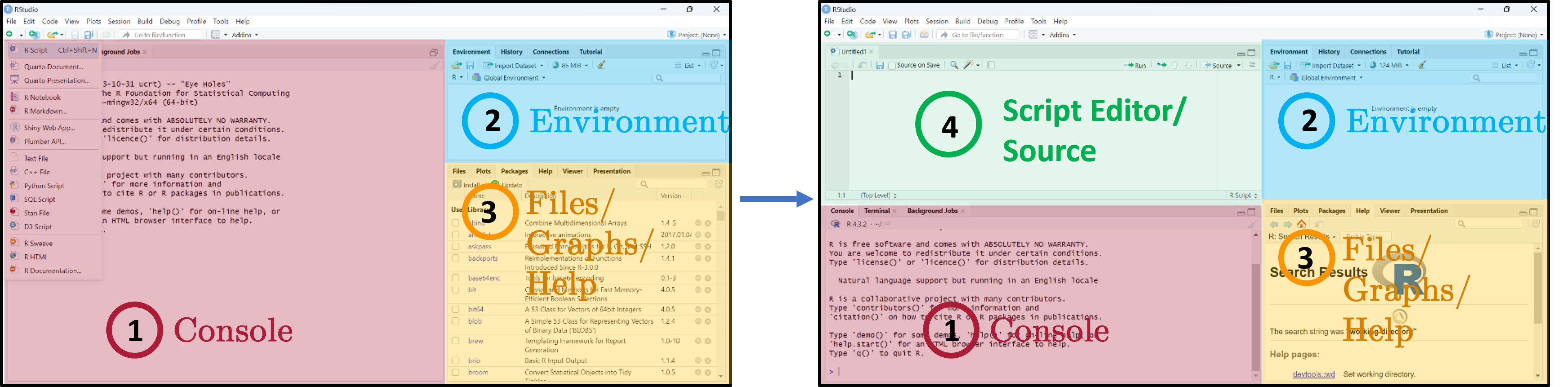
RStudio interface
Three main panes when you open RStudio:
- Console — where R code is executed.
- Global Environment / History — your in-memory objects.
- Files / Plots / Packages / Help / Viewer — the swiss-army-knife pane.
- Script Editor — where you write and save your R scripts.
Writing code in RStudio
- Load and save
.Rscripts. - Keeps a record of your analysis — show others how it was run, repeat later.
- Edit code without accidentally running it.
- Where R code is executed.
- You can also run code interactively here, but it is NOT saved on disk.
- See output and errors directly.
Write and run code through the Script Editor. Use the Console only for quick exploration.
Setting the working directory
- The working directory (
wd) is where R looks for and saves files. - Two ways to set it: via the menu, or in code with
setwd(). - Always verify with
getwd()to avoid confusion about where your files are being read from or written to. - Tip: set the working directory at the start of your script so it’s clear where files are coming from and going to. Avoid hardcoding paths if you plan to share your code with others or run it on different machines.
Getting started — running code
The blinking cursor in the Script Editor prompts you to write. The Console shows > when R is ready.
Four ways to run code:
- Ctrl/Cmd + Enter — run current line / selection.
- Click Run.
- Ctrl + Shift + Enter — run all lines in the editor.
- Highlight lines and click Run.
Short exercise
Type 1 + 2*8 and log(10). Result appears in the Console with a leading [1] (the index of the first element on that line).
What are packages?
- Packages expand what you can do beyond base R.
- A collection of functions, data sets, and other R objects under one name.
- Install from repositories: CRAN, GitHub, etc.
- The Packages tab lists installed packages; click Update to upgrade; the small × uninstalls.
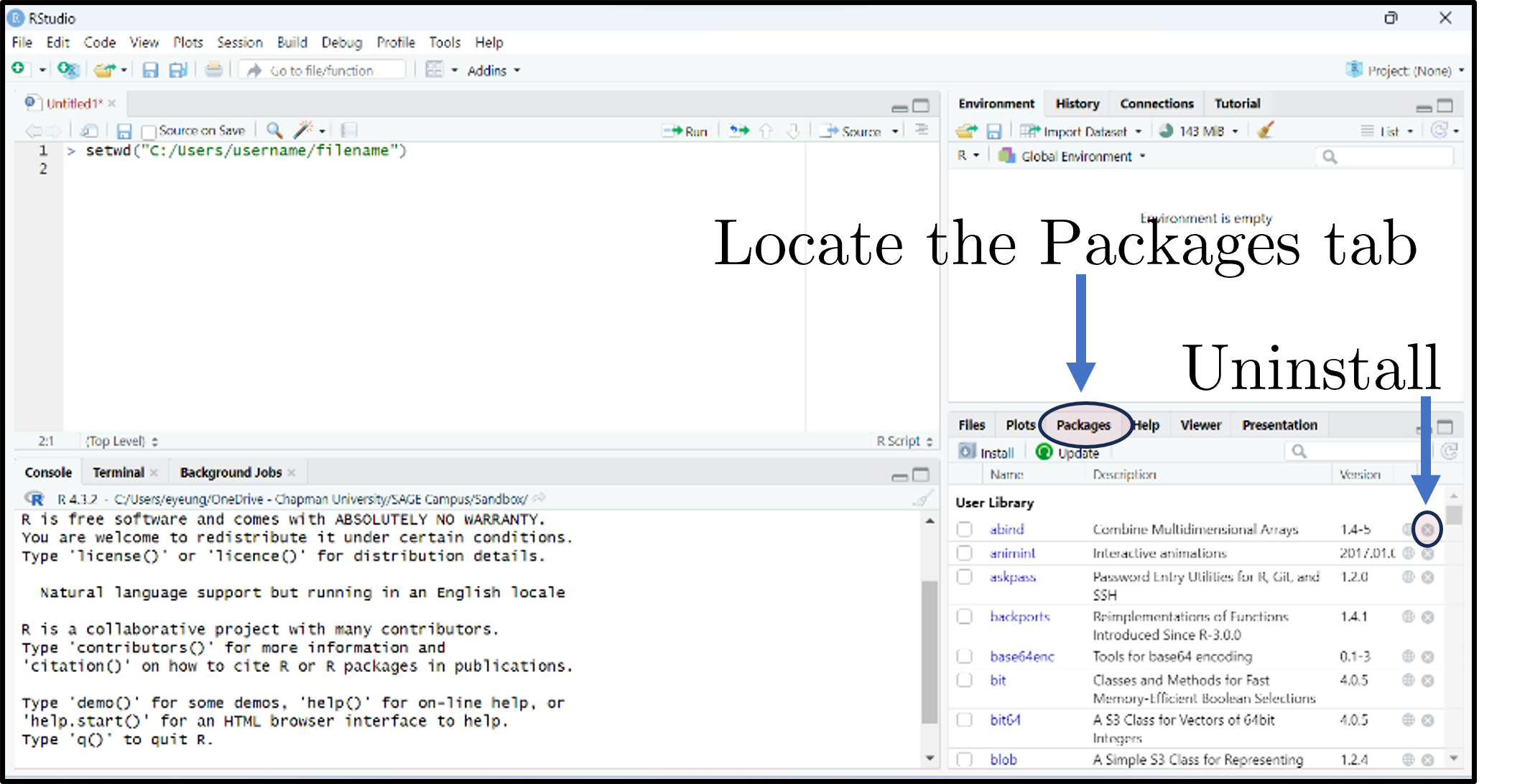
Installing and loading packages
Or click Install in the Packages tab.
Or tick the package’s checkbox in the Packages tab.
Short exercise
Install and load the tidyverse, tidyquant, and Quandl packages.
1.3 Live Coding Session 1
- 1.1 Course objectives
- 1.2 Introduction to R
- 1.3 Live Coding Session 1
- 1.4 Conclusion of Lecture 1
- Creating objects / variables
- Four data types in R
- Variables and basic operations
- Five basic data structures
- Vectors
- Matrices
- Data frames
- Lists
- Functions
- Loops
- Data import & export — file types
- Data import & export — examples
- Saving your script
- Closing an R session
Creating objects / variables
- The “<-” operator assigns the value on the right to the name on the left.
- Avoid long object names, be descriptive
- Don’t reuse names of existing R functions
- No blank spaces - use
_or. - R is case-sensitive (
xandXare different) - Use
=for function arguments, but “<-” for assignment to avoid confusion - Don’t use
=for assignment in scripts, as it can lead to bugs and readability issues
Four data types in R
- Numeric — numbers that may contain decimals.
- Integer — special case, no decimals (suffix
L).
- Integer — special case, no decimals (suffix
- Character — text (strings); wrap in
" ". - Factor — special character used for categorical data (e.g., male/female, months).
- Logical — Boolean (
TRUE/FALSE).
class(objectname)returns the higher-level label R uses to decide which functions and behaviors apply ("data.frame","factor","matrix","numeric").
Variables and basic operations
- Assign and compute financial metrics — returns, prices, ratios — quickly during empirical work
- Use factors to manage categorical variables like sectors, credit ratings, or time periods
- Logical variables are useful for filtering data frames (e.g.,
subset(myData, myData$sector == "Tech"))
Five basic data structures
- Individual values — single scalars.
- Vectors — ordered collections of elements (one type only).
- Matrices — 2D arrays; rows × columns; one type only.
- Data frames — like spreadsheets; columns can have different types.
- Lists — flexible containers that can hold vectors, matrices, data frames, even other lists.
Vectors
# Create
myVector <- c(3, 4, -1.1, pi)
# Inspect / aggregate
myVector[3] # third element: -1.1
length(myVector) # 4
sum(myVector) # ~9.04
mean(myVector) # ~2.26
var(myVector) # ~5.21
sort(myVector) # ascending: -1.1, 3.0, 3.14, 4.0
quantile(myVector, c(0.1, 0.25, 0.95))
# Vectorized arithmetic — no loops needed
myVector + 2
myVector * myVector
# Coercion: mixed types collapse to character
mixed <- c(1, "two", TRUE) # "1" "two" "TRUE"
# Named vectors
namedVec <- c(apple = 5, banana = 3)
namedVec["banana"] # 3- Basic syntax:
c()combines values into a vector; indexing with[]retrieves elements - Vectors are the building blocks of more complex structures. They allow you to store and manipulate sequences of numbers, text, or logical values efficiently
- Vectorized operations enable you to perform calculations on entire vectors without writing explicit loops, which is faster and more concise
- Named vectors can improve readability and allow for easier access to specific elements by name rather than index
Matrices
myMatrix <- matrix(c(2, 1, 0, -9, 5, 0), nrow = 2, byrow = TRUE)
dim(myMatrix) # [1] 2 3
myMatrix[1, 1] <- 2.1415 # modify cell
myMatrix[, 2] # second column
# Linear algebra
anotherMatrix <- matrix(c(5, -1, 7, 0, 2, -1), nrow = 2, byrow = TRUE)
transposedMatrix <- t(anotherMatrix)
matrixProduct <- myMatrix %*% transposedMatrix # 2x2 product
matrixInverse <- solve(matrixProduct) # inverse
# Aggregations
rowSums(myMatrix)
colMeans(myMatrix)
# Bind: combine rows / cols
cbind(myMatrix, c(6, 7))- Basic syntax:
matrix()creates a matrix from a vector of values; specifynrowandncolto set dimensions;byrow = TRUEfills by rows instead of columns - Matrices are 2D structures that can only hold one type of data (e.g., all numeric)
- They are essential for linear algebra operations, which are common in finance1
- Functions like
rowSums(),colMeans(), andcbind()allow you to perform common matrix manipulations without writing loops
Data frames
student_name <- c("Student A", "Student B", "Student C")
grade <- c(1.7, 2.3, 5.0)
students <- data.frame(student_name, grade)
students$grade # column access
colnames(students)[2] <- "exam2021"
students[2:3, ] # rows 2-3
# Add columns
students$degree <- c("Bachelor", "Master", "Bachelor")
students$pass_fail <- c("Pass", "Pass", "Fail")
table(students$degree, students$pass_fail)
# Summary
summary(students)
# Subset by condition
subset(students, exam2021 > 2)
# Add row
newStudent <- data.frame(student_name = "Student D", exam2021 = 3.5,
degree = "PhD", pass_fail = "Pass")
students <- rbind(students, newStudent)
# Merge with another frame
ages <- data.frame(student_name = c("Student A", "Student B", "Student C"),
age = c(20, 22, 21))
merged <- merge(students, ages, by = "student_name")- Basic syntax:
data.frame()creates a data frame from vectors; columns can have different types; access with$or[] - Data frames are the most common structure for tabular data in R. They can hold different types of data in different columns (e.g., numeric grades, character names, factors for degree)
- Adding new columns is straightforward, and you can easily summarize or subset the data
- Merging data frames is common when you have related datasets1 that share a key
Lists
scoresList <- list(
scores = c(95, 85, 92),
names = c("Alice", "Bob", "Carol"),
passed = c(TRUE, TRUE, FALSE)
)
str(scoresList)
scoresList$scores[3] <- 90 # update Carol's score
scoresList$comments <- "All passed" # add element
# Nested list
nestedList <- list(course = "Math 101", details = scoresList)
nestedList$details$names[1] # "Alice"
# Convert to data frame (elements need equal length)
scoresDF <- as.data.frame(scoresList)- Basic syntax:
list()creates a list; access with$or[]; lists can contain any type of object, including other lists - Lists are flexible containers that can hold a variety of objects, making them useful for storing complex data structures or results from functions that return multiple outputs
- They are particularly useful when you want to return multiple related objects from a function without having to combine them into a single data frame or matrix
Functions
# Define
bmiCalc <- function(height, weight) {
bmi <- weight / (height ^ 2)
return(round(bmi, 1))
}
# Call
myBmi <- bmiCalc(1.75, 70) # 22.9
# Default arguments
bmiCalcDefault <- function(height, weight = 70) {
round(weight / (height ^ 2), 1)
}
bmiCalcDefault(1.75) # 22.9
# Multiple returns via list
bmiAdvanced <- function(height, weight) {
bmi <- weight / (height ^ 2)
category <- ifelse(bmi < 18.5, "Underweight",
ifelse(bmi < 25, "Normal", "Overweight"))
list(bmi = round(bmi, 1), category = category)
}
result <- bmiAdvanced(1.75, 70)
result$bmi # 22.9
result$category # "Normal"- Basic syntax:
function(arg1, arg2) { ... }defines a function; usereturn()to specify output; functions can have default arguments and return multiple values via lists - Functions allow you to encapsulate reusable code, making your scripts cleaner and more modular
- Functions are essential for performing repeated calculations, especially when you need to apply the same logic to different inputs
Loops
- Basic syntax:
for (var in sequence) { ... }iterates over elements;while (condition) { ... }continues until condition is false - Loops are useful for certain tasks, but in R, they can be inefficient for data manipulation. Whenever possible, use vectorized operations or apply functions instead of loops for better performance
- In practice, you’ll often use functions from packages like
dplyrthat abstract away the need for explicit loops when working with data frames
Data import & export — file types
You will rarely build matrices/data frames by hand. Common ways to read data:
- Text —
.TXT(readLines()) - Tabular —
.CSV,.TSV(read.table(),readr::read_csv()) - Excel —
.XLSX(xlsx,readxl) - Google sheets —
googlesheets4 - Statistics programs — SPSS, SAS (
haven) - Databases — MySQL (
RMySQL)
Data import & export — examples
# CSV
write.csv(objectname, file = "output_file.csv", row.names = FALSE)
# Tab-delimited
write.table(objectname, file = "output_file.txt",
sep = "\t", row.names = FALSE, col.names = TRUE)
# Excel
library(writexl)
write_xlsx(objectname, "output_file.xlsx")
# RDS
saveRDS(objectname, file = "output_file.rds")The argument
sep =may not be necessary — R recognises,in.csvand whitespace in.txt. But sometimes R won’t read correctly unless you specify it
Saving your script
- Save every step in an
.Rscript. Comment lines start with#and are ignored at run time. - File → Save, or click the disk icon, or Ctrl/Cmd + S.
- Unsaved scripts (or unsaved edits) show red text and an asterisk in the tab name.
Closing an R session
Three ways to close RStudio:
- File → Quit Session
- The red ✕ in the top-right corner.
- Run
q()in the Console.
Don’t save the workspace image (.RData)
Unless (a) you ran something very expensive, or (b) you’re nearly done with the project. Starting clean each session avoids hidden state from previous runs.
1.4 Conclusion of Lecture 1
- 1.1 Course objectives
- 1.2 Introduction to R
- 1.3 Live Coding Session 1
- 1.4 Conclusion of Lecture 1
- Course at a glance
- Further reading
- Prepare before next lecture
- See you next time
- References
Course at a glance
Basics
Week 1
29.10.2025
Course objectives, schedule, assignments · Introduction to R · Live coding
- Course objectives, schedule and assignments
- Introduction to R and RStudio
- Live coding: variables, vectors, matrices, data frames, lists, functions, loops
- Data import and export
Data Handling & Visualization
Week 2
05.11.2025
API access, merging, cleansing, transforming and visualising financial data in R · Introduction to Overleaf
- API access (Nasdaq Data Link / Quandl, FRED, Yahoo, Coingecko, Polygon)
- Import and cleanse: read_csv, mutate, types
- Merge and append data (merge, bind_rows)
- Filter and mutate (dplyr): subset rows, derive variables
- Group by and summarise
- Pivot wide / long
- Data visualization with ggplot2 (six-step pipeline)
- Introduction to LaTeX and Overleaf
Statistical Analysis
Week 3
12.11.2025
Descriptive · inferential · modelling — applied in R
- Descriptive statistics in R
- Correlation matrix and Pearson correlation test
- t-Test and Wilcoxon test
- Shapiro-Wilk and Kolmogorov-Smirnov tests
- Linear regression with fixed effects
- Clustered standard errors
- Exporting regression tables with stargazer
- Discussion of Assignment I (Problem Set)
Academic Publishing & Refereeing
Week 4
19.11.2025
What makes a great empirical paper · publication process · how to write a referee report
- What makes a good empirical paper (contribution, identification, write-up)
- The publication process step by step
- Top finance and economics journals
- Bad outcome vs revise & resubmit
- Referee Reports — summary, major issues, minor issues
- Referee checklist (question, identification, data, econometrics, results)
- Discussion of Assignment II (Referee Report)
Brown Bag Seminar
Week 13
20.01.2026
Engage with doctoral research and prepare your referee report
- Doctoral research presentations
- Apply empirical / writing tips for the referee report
- Group discussion and Q&A
Further reading
- James et al. (2021) — free PDF and exercises at https://www.statlearning.com/.
- Scheuch, Voigt, and Weiss (2023) — finance-specific tidyverse cookbook, free at https://www.tidy-finance.org/r/.
- Wickham and Çetinkaya-Rundel (2023) — second edition, free at https://r4ds.hadley.nz/.
- Kaggle Discussions (2021) — community-curated list of additional free R books.
Prepare before next lecture
- Document today’s code in a clean way and save as
.Rmd. - Set up the Nasdaq Data Link API:
- Register a free account at https://data.nasdaq.com/ (formerly Quandl).
- Log in, go to your profile, copy your personal API key.
- Optional challenge: load the
Quandllibrary, set your key withQuandl.api_key("your_key_here"), and test loading data. We’ll walk through this in Lecture 2.
See you next time
Reminder
- Register for “exam” 13337 in
campusonlineby 30 November 2025. - Bring your laptop with R + RStudio installed and the
tidyverse,tidyquant,Quandlpackages already loaded. - Lecture 2: R Data Handling and Visualization — APIs, merging, cleansing,
dplyr,ggplot2.
References
Institute of Strategic Management and Finance · Ulm University